Data frame Manipulation with dplyr
Last updated on 2024-08-19 | Edit this page
Estimated time: 40 minutes
Overview
Questions
- How can I manipulate dataframes without repeating myself?
Objectives
- To be able to use the six main dataframe manipulation ‘verbs’ with
pipes in
dplyr. - To understand how
group_by()andsummarize()can be combined to summarize datasets. - Be able to analyze a subset of data using logical filtering.
Manipulation of dataframes means many things to many researchers, we often select certain observations (rows) or variables (columns), we often group the data by a certain variable(s), or we even calculate summary statistics. We can do these operations using the normal base R operations:
R
gapminder <- read.csv("data/gapminder_data.csv", header = TRUE)
mean(gapminder[gapminder$continent == "Africa", "gdpPercap"])
OUTPUT
[1] 2193.755R
mean(gapminder[gapminder$continent == "Americas", "gdpPercap"])
OUTPUT
[1] 7136.11R
mean(gapminder[gapminder$continent == "Asia", "gdpPercap"])
OUTPUT
[1] 7902.15But this isn’t very efficient, and can become tedious quickly because there is a fair bit of repetition. Repeating yourself will cost you time, both now and later, and potentially introduce some nasty bugs.
The dplyr package
- Introduce the
dplyrpackage as a simpler, more intuitive way of doing subsetting. - Unlike other SWC and DC R lessons, this lesson does
not include data reshaping with
tidyras it isn’t used in the rest of the workshop.
Luckily, the dplyr package
provides a number of very useful functions for manipulating dataframes
in a way that will reduce the above repetition, reduce the probability
of making errors, and probably even save you some typing. As an added
bonus, you might even find the dplyr grammar easier to
read.
Here we’re going to cover 6 of the most commonly used functions as
well as using pipes (%>%) to combine them.
select()filter()group_by()summarize()-
count()andn() mutate()
If you have have not installed this package earlier, please do so:
R
install.packages('dplyr')
Now let’s load the package:
R
library("dplyr")
Using select()
If, for example, we wanted to move forward with only a few of the
variables in our dataframe we could use the select()
function. This will keep only the variables you select.
R
year_country_gdp <- select(gapminder, year, country, gdpPercap)
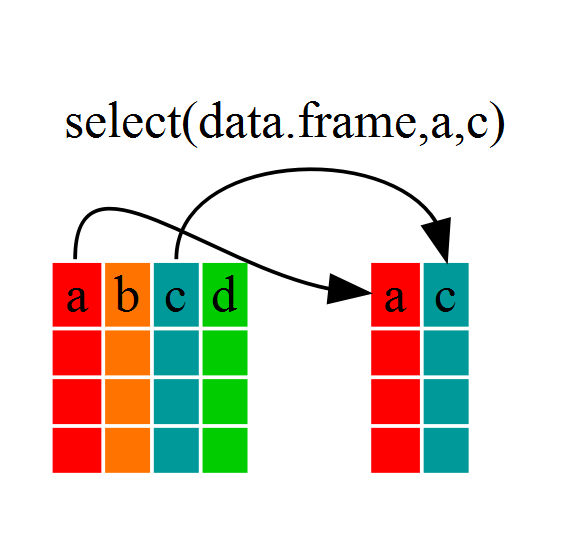
If we open up year_country_gdp we’ll see that it only
contains the year, country and gdpPercap. Above we used ‘normal’
grammar, but the strengths of dplyr lie in combining
several functions using pipes. Since the pipes grammar is unlike
anything we’ve seen in R before, let’s repeat what we’ve done above
using pipes.
R
year_country_gdp <- gapminder %>% select(year,country,gdpPercap)
To help you understand why we wrote that in that way, let’s walk
through it step by step. First we summon the gapminder data
frame and pass it on, using the pipe symbol %>%, to the
next step, which is the select() function. In this case we
don’t specify which data object we use in the select()
function since in gets that from the previous pipe. Fun
Fact: You may have encountered pipes before in the shell. In R,
a pipe symbol is %>% while in the shell it is
| but the concept is the same!
Using filter()
If we now wanted to move forward with the above, but only with
European countries, we can combine select and
filter
R
year_country_gdp_euro <- gapminder %>%
filter(continent == "Europe") %>%
select(year, country, gdpPercap)
R
year_country_lifeExp_Africa <- gapminder %>%
filter(continent=="Africa") %>%
select(year,country,lifeExp)
As with last time, first we pass the gapminder dataframe to the
filter() function, then we pass the filtered version of the
gapminder data frame to the select() function.
Note: The order of operations is very important in this
case. If we used ‘select’ first, filter would not be able to find the
variable continent since we would have removed it in the previous
step.
Using group_by() and summarize()
Now, we were supposed to be reducing the error prone repetitiveness
of what can be done with base R, but up to now we haven’t done that
since we would have to repeat the above for each continent. Instead of
filter(), which will only pass observations that meet your
criteria (in the above: continent=="Europe"), we can use
group_by(), which will essentially use every unique
criteria that you could have used in filter.
R
str(gapminder)
OUTPUT
'data.frame': 1704 obs. of 6 variables:
$ country : chr "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num 779 821 853 836 740 ...R
gapminder %>% group_by(continent) %>% str()
OUTPUT
gropd_df [1,704 × 6] (S3: grouped_df/tbl_df/tbl/data.frame)
$ country : chr [1:1704] "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int [1:1704] 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num [1:1704] 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr [1:1704] "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num [1:1704] 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num [1:1704] 779 821 853 836 740 ...
- attr(*, "groups")= tibble [5 × 2] (S3: tbl_df/tbl/data.frame)
..$ continent: chr [1:5] "Africa" "Americas" "Asia" "Europe" ...
..$ .rows : list<int> [1:5]
.. ..$ : int [1:624] 25 26 27 28 29 30 31 32 33 34 ...
.. ..$ : int [1:300] 49 50 51 52 53 54 55 56 57 58 ...
.. ..$ : int [1:396] 1 2 3 4 5 6 7 8 9 10 ...
.. ..$ : int [1:360] 13 14 15 16 17 18 19 20 21 22 ...
.. ..$ : int [1:24] 61 62 63 64 65 66 67 68 69 70 ...
.. ..@ ptype: int(0)
..- attr(*, ".drop")= logi TRUEYou will notice that the structure of the dataframe where we used
group_by() (grouped_df) is not the same as the
original gapminder (data.frame). A
grouped_df can be thought of as a list where
each item in the listis a data.frame which
contains only the rows that correspond to the a particular value
continent (at least in the example above).
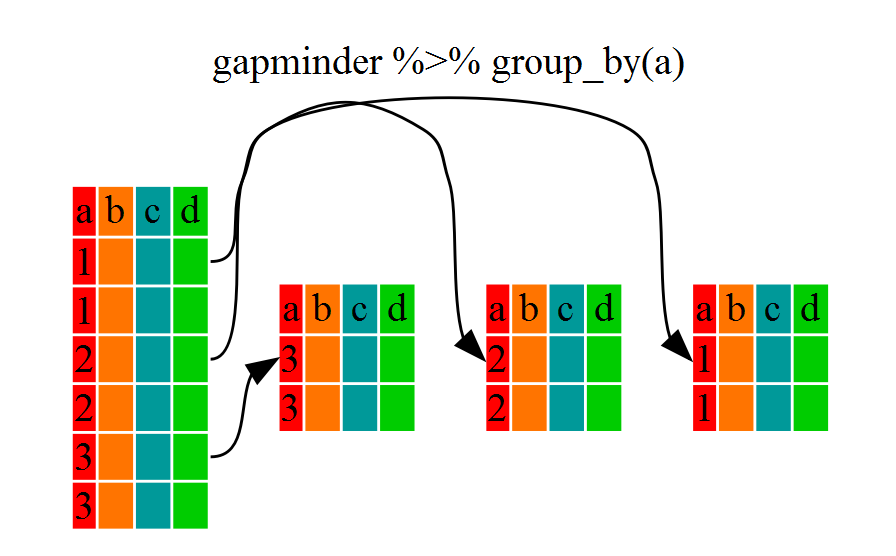
Using summarize()
The above was a bit on the uneventful side but
group_by() is much more exciting in conjunction with
summarize(). This will allow us to create new variable(s)
by using functions that repeat for each of the continent-specific data
frames. That is to say, using the group_by() function, we
split our original dataframe into multiple pieces, then we can run
functions (e.g. mean() or sd()) within
summarize(). To end our group_by(), we need to
close off the pipe by using ungroup().
R
gdp_bycontinents <- gapminder %>%
group_by(continent) %>%
summarize(mean_gdpPercap = mean(gdpPercap)) %>%
ungroup()
gdp_bycontinents
OUTPUT
# A tibble: 5 × 2
continent mean_gdpPercap
<chr> <dbl>
1 Africa 2194.
2 Americas 7136.
3 Asia 7902.
4 Europe 14469.
5 Oceania 18622.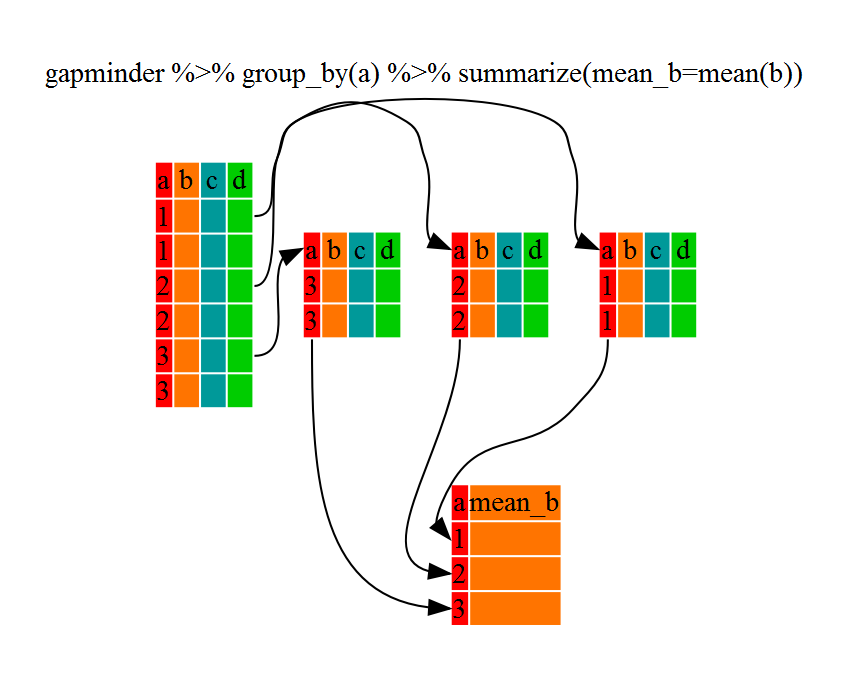
That allowed us to calculate the mean gdpPercap for each continent, but it gets even better.
R
lifeExp_bycountry <- gapminder %>%
group_by(country) %>%
summarize(mean_lifeExp=mean(lifeExp)) %>%
ungroup()
lifeExp_bycountry %>%
filter(mean_lifeExp == min(mean_lifeExp) | mean_lifeExp == max(mean_lifeExp))
OUTPUT
# A tibble: 2 × 2
country mean_lifeExp
<chr> <dbl>
1 Iceland 76.5
2 Sierra Leone 36.8Another way to do this is to use the dplyr function
arrange(), which arranges the rows in a data frame
according to the order of one or more variables from the data frame. It
has similar syntax to other functions from the dplyr
package. You can use desc() inside arrange()
to sort in descending order.
R
lifeExp_bycountry %>%
arrange(mean_lifeExp) %>%
head(1)
OUTPUT
# A tibble: 1 × 2
country mean_lifeExp
<chr> <dbl>
1 Sierra Leone 36.8R
lifeExp_bycountry %>%
arrange(desc(mean_lifeExp)) %>%
head(1)
OUTPUT
# A tibble: 1 × 2
country mean_lifeExp
<chr> <dbl>
1 Iceland 76.5The function group_by() allows us to group by multiple
variables. Let’s group by year and
continent.
R
gdp_bycontinents_byyear <- gapminder %>%
group_by(continent, year) %>%
summarize(mean_gdpPercap = mean(gdpPercap)) %>%
ungroup()
That is already quite powerful, but it gets even better! You’re not
limited to defining 1 new variable in summarize().
R
gdp_pop_bycontinents_byyear <- gapminder %>%
group_by(continent,year) %>%
summarize(mean_gdpPercap = mean(gdpPercap),
sd_gdpPercap = sd(gdpPercap),
mean_pop = mean(pop),
sd_pop = sd(pop)) %>%
ungroup()
count() and n()
A very common operation is to count the number of observations for
each group. The dplyr package comes with two related
functions that help with this.
For instance, if we wanted to check the number of countries included
in the dataset for the year 2002, we can use the count()
function. It takes the name of one or more columns that contain the
groups we are interested in, and we can optionally sort the results in
descending order by adding sort=TRUE:
R
gapminder %>%
filter(year == 2002) %>%
count(continent, sort = TRUE)
OUTPUT
continent n
1 Africa 52
2 Asia 33
3 Europe 30
4 Americas 25
5 Oceania 2If we need to use the number of observations in calculations, the
n() function is useful. For instance, if we wanted to get
the standard error of the life expectancy per continent:
R
gapminder %>%
group_by(continent) %>%
summarize(sd_le = sd(lifeExp), # standard deviation
n_le = n(), # number of observations
se_le = sd_le/sqrt(n_le)) %>% # standard error
ungroup()
OUTPUT
# A tibble: 5 × 4
continent sd_le n_le se_le
<chr> <dbl> <int> <dbl>
1 Africa 9.15 624 0.366
2 Americas 9.35 300 0.540
3 Asia 11.9 396 0.596
4 Europe 5.43 360 0.286
5 Oceania 3.80 24 0.775In the above example, we chained together several summary operations
to simplify our calculation of standard error. We can also chain
operations to calculate the minimum, maximum,
mean and se of each continent’s per-country
life-expectancy. This time, we will calculate the se in
just one line:
R
gapminder %>%
group_by(continent) %>%
summarize(
mean_le = mean(lifeExp),
min_le = min(lifeExp),
max_le = max(lifeExp),
se_le = sd(lifeExp)/sqrt(n())) %>%
ungroup()
OUTPUT
# A tibble: 5 × 5
continent mean_le min_le max_le se_le
<chr> <dbl> <dbl> <dbl> <dbl>
1 Africa 48.9 23.6 76.4 0.366
2 Americas 64.7 37.6 80.7 0.540
3 Asia 60.1 28.8 82.6 0.596
4 Europe 71.9 43.6 81.8 0.286
5 Oceania 74.3 69.1 81.2 0.775Using mutate()
We can also create new variables prior to (or even after) summarizing
information using mutate().
R
gdp_pop_billion <- gapminder %>%
mutate(gdp_billion = gdpPercap*pop/10^9)
head(gdp_pop_billion)
OUTPUT
country year pop continent lifeExp gdpPercap gdp_billion
1 Afghanistan 1952 8425333 Asia 28.801 779.4453 6.567086
2 Afghanistan 1957 9240934 Asia 30.332 820.8530 7.585449
3 Afghanistan 1962 10267083 Asia 31.997 853.1007 8.758856
4 Afghanistan 1967 11537966 Asia 34.020 836.1971 9.648014
5 Afghanistan 1972 13079460 Asia 36.088 739.9811 9.678553
6 Afghanistan 1977 14880372 Asia 38.438 786.1134 11.697659Other great resources
- R for Data Science
- Data Wrangling Cheat sheet
- Introduction to dplyr
- Data wrangling with R and RStudio
Key Points
- Use the
dplyrpackage to manipulate dataframes. - Use
select()to choose variables from a dataframe. - Use
filter()to choose data based on values. - Use
group_by()andsummarize()to work with subsets of data. - Use
count()andn()to obtain the number of observations in columns. - Use
mutate()to create new variables.
